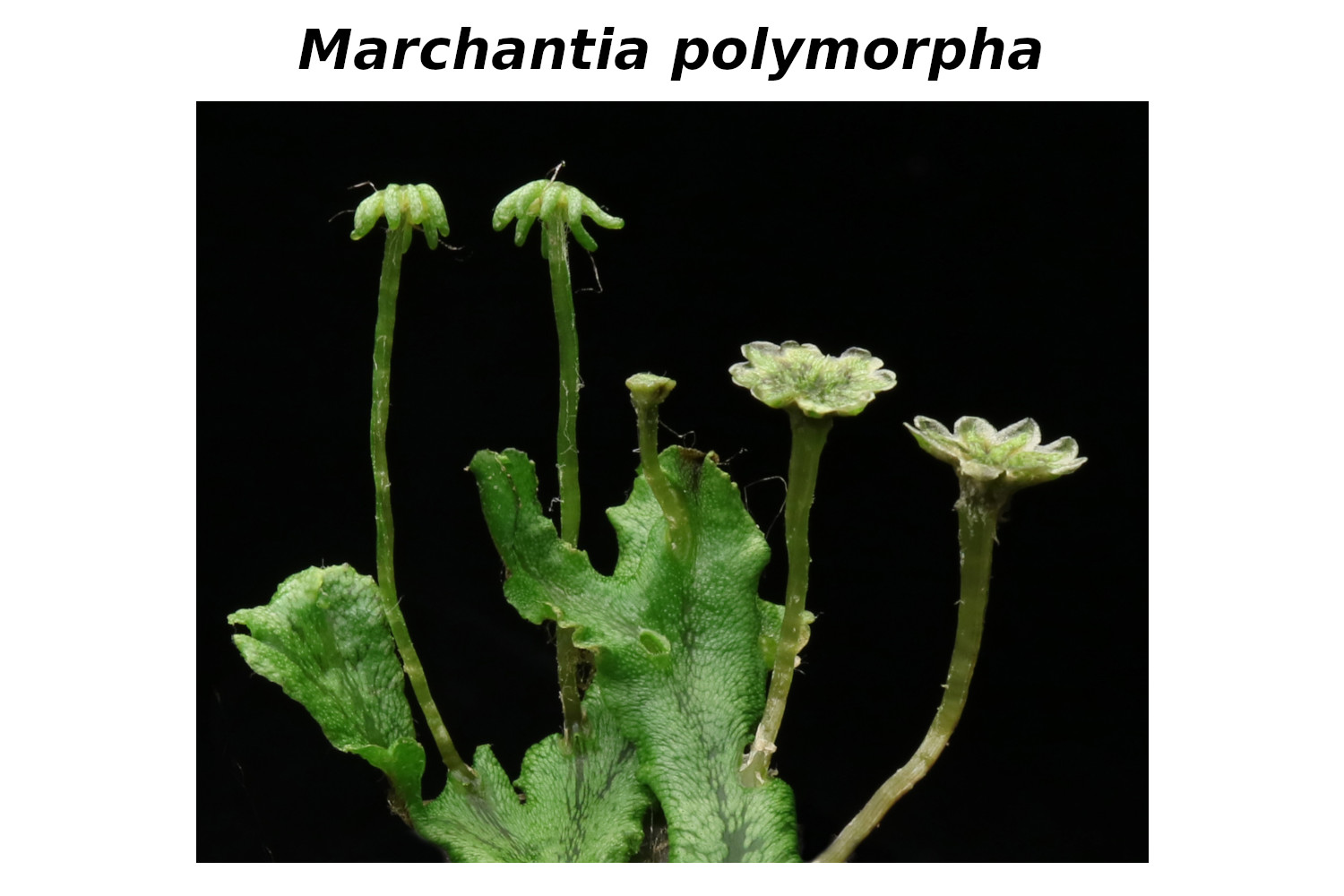
Our research team analyses molecular mechanisms that contribute to generate the enormous diversity of land plants. These processes are key to understand how our extant land plant species adapted to extremely variable conditions such that they can grow and propagate in almost all climate zones. We investigate transcription factors that control plant development and thereby shape plant morphogenesis. We conduct comparative research approaches using classical plant model organisms, such as Arabidopsis thaliana together with other evolutionary informative angiosperm and basal land plant species. Current research focuses on the liverwort Marchantia polymorpha to unravel the role of redox-processes in developmental processes and for an adaptation to a terrestrial plant life style.

We apply a combination of diverse genetic (genome editing), biochemical (redox-EMSAs, Y2H), transcriptome (RNAseq), microscopy (FRET-FLIM, superresolution) and expression (whole mount/in situ mRNA, fluorescent-reporters) studies.
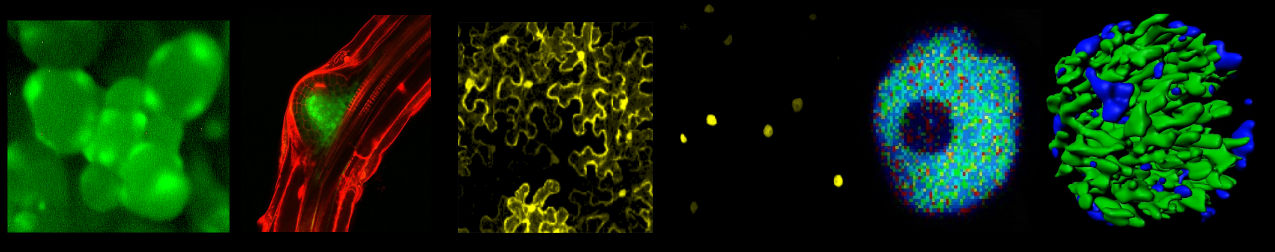



Click on the image for further information
Zachgo S. (2023) Nuclear redox processes in land plant development and stress adaptation. Biol. Chem., DOI 10.1515/hsz-2022-0288.
Bowman J., Arteaga-Vazquez M., Berger F., et al., Zachgo S. (2022) The renaissance and enlightenment of Marchantia as a model system. Plant Cell 34, 3512–3542. DOI 10.1093/plcell/koac219.
Althoff F., Wegner L., Ehlers K., Buschmann H., Zachgo S. (2022) Developmental plasticity of the amphibious liverwort Riccia fluitans. Front. Plant Sci., DOI 10.3389/fpls.2022.909327.
Maß L., Holtmannspötter M., Zachgo S. (2020) Dual-color 3D-dSTORM colocalization and quantification of ROXY1 and RNAPII variants throughout the transcription cycle in root meristem nuclei. The Plant Journal, DOI 10.111/tpj.14986.
Althoff F., Zachgo S. (2020) Transformation of Riccia fluitans, an amphibious liverwort dynamically responding to environmental changes. International Journal of Molecular Sciences 21(15), 5410. DOI.org/10.3390/ijms21155410.
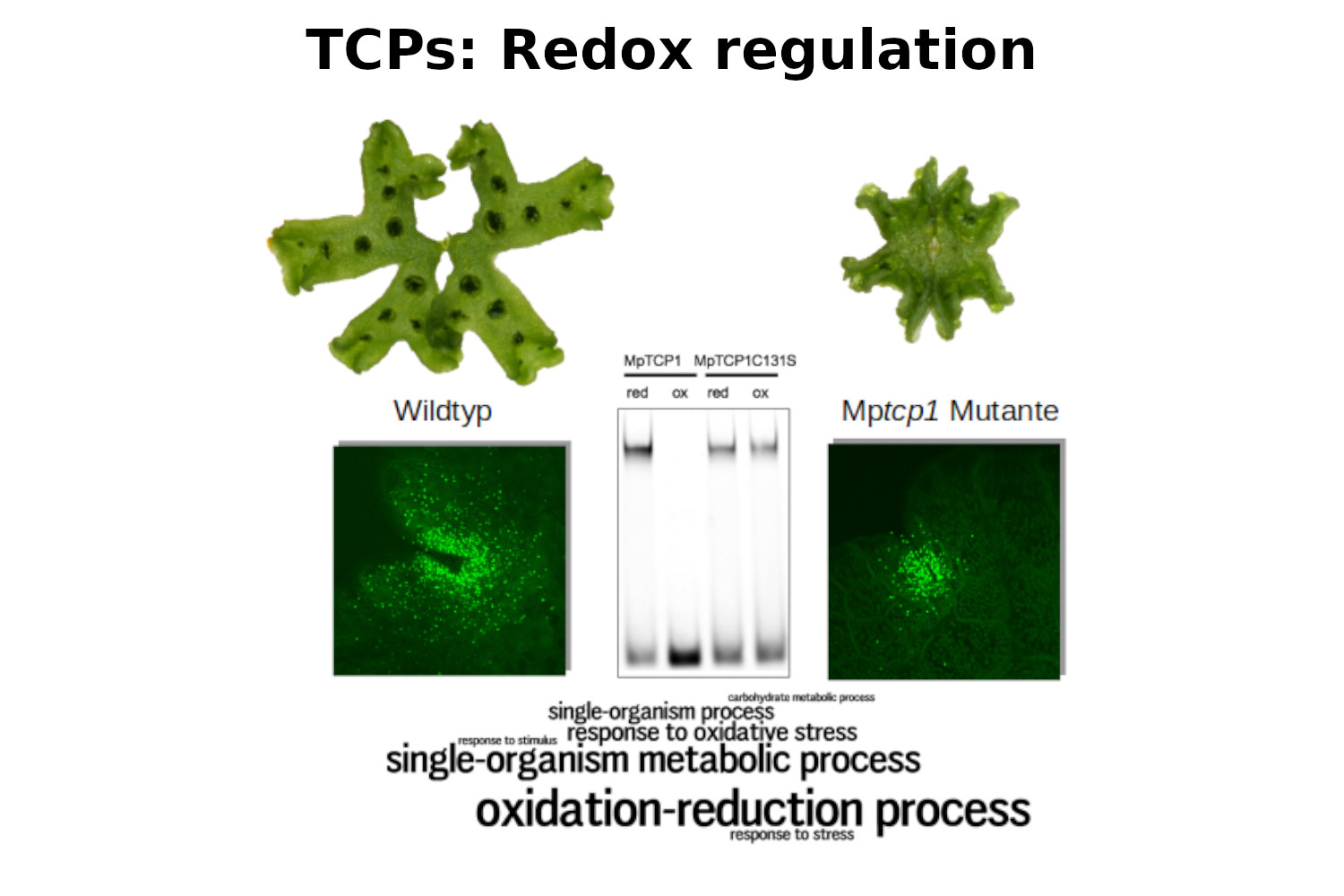
Busch A., Deckena M., Almeida-Trapp M., Kopischke S., Kock C., Schuessler E., Tsiantis M., Mithofer A., Zachgo S. (2019) MpTCP1 controls cell proliferation and redox processes in Marchantia polymorpha. New Phytologist, DOI 10.1111/nph.16132.
Nishiyama T., et al. (Chara genome consortium), Zachgo S., Langdale J., Maumus F., Van Der Straeten D., Gould S.B., Rensing S.A. (2018) The Chara genome: secondary complexity and implications for plant terrestrialization. Cell 174, 448-464.
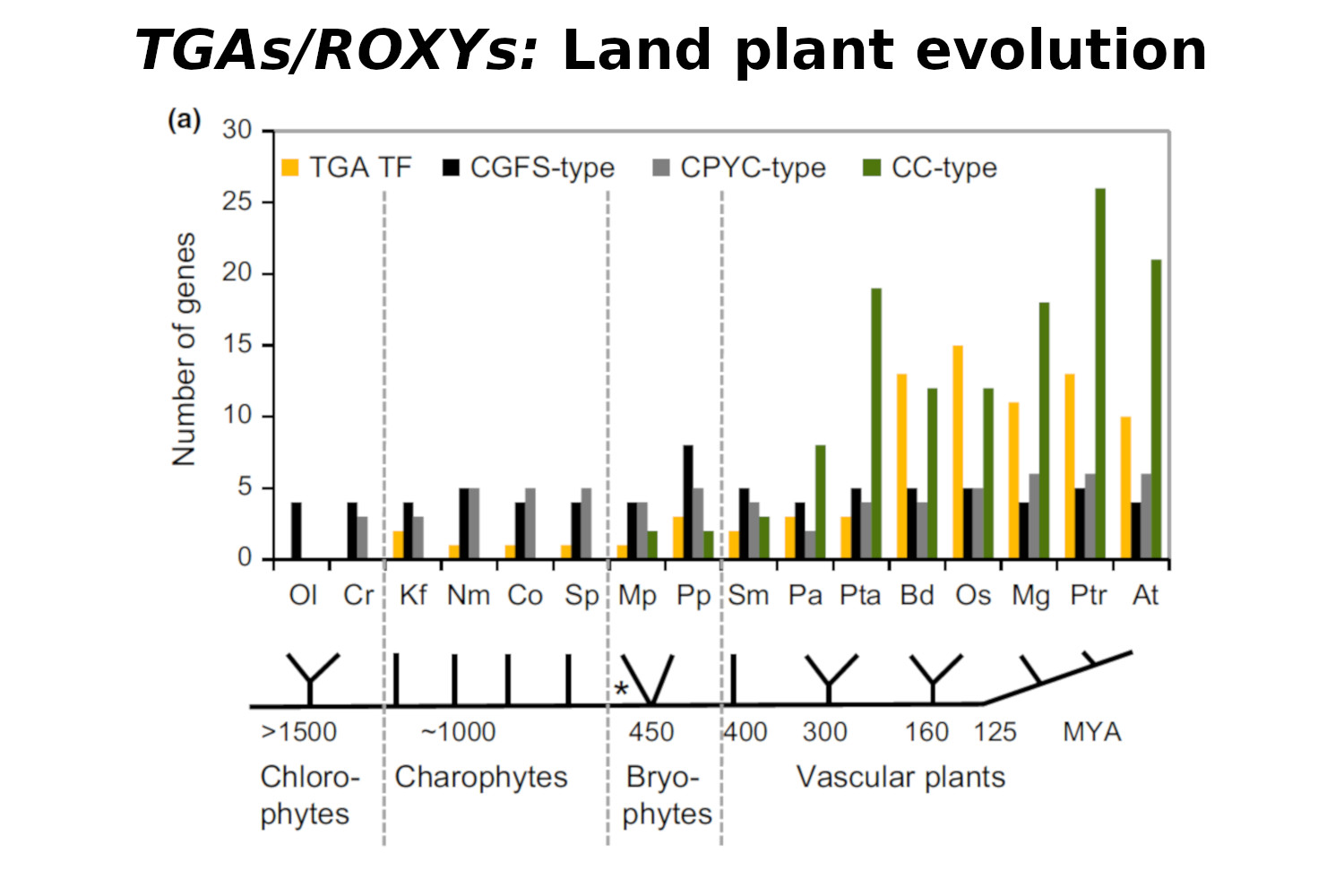
Gutsche N., Holtmannspötter M., Maß L., O’Donoghue M., Busch A., Lauri A., Schubert V., Zachgo S. (2017) Conserved redox-dependent DNA binding of ROXY glutaredoxins with TGA transcription factors. Plant Direct, DOI 10.1002/pld3.30.
Bowman J.L. et al. (Marchantia genome consortium), Zachgo S., Schmutz J. (2017) Insights into Land Plant Evolution Garnered from the Marchantia polymorpha Genome. Cell 171, 287-304.
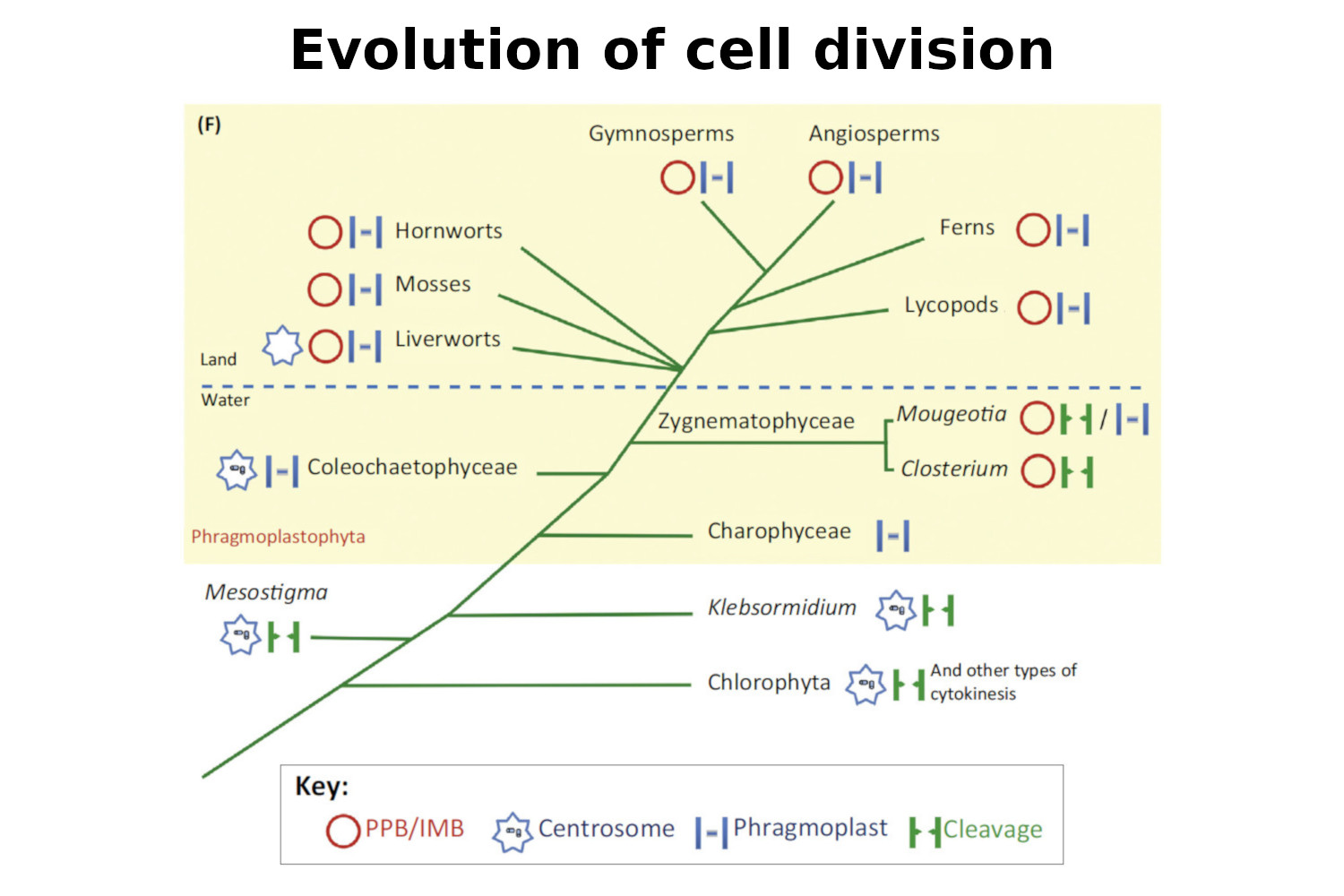
Buschmann H., Zachgo S. (2016) The evolution of cell division: from streptophyte algae to land plants. Trends in Plant Science, 21, 872-883.
Borgmann P., Oevermann S., Zachgo S. (2023) Hotspotuntersuchungen nationaler Verantwortungsarten der Gefäßpflanzen in Nordrhein-Westfalen und Schleswig-Holstein. Natur und Landschaft 98, Heft 5.
Borgmann P., Burkart M., Lauterbach D., Listl D., Martens A., Nick P., Oevermann S., Poschlod P., Radkowitsch A., Reisch C., Stevens A.-D., Straubinger C., Zippel E., Zachgo S. (2015) WIPs-De: Wildpflanzenschutz Deutschland - Ein Projekt des Bundesprogramms zur Biologischen Vielfalt. Natur und Landschaft 90, Heft 12, S. 550-555.
Borgmann P., Oevermann S., Friesen N. und Zachgo S. (2014) Die Genbank für Wildpflanzen für Ernährung und Landwirtschaft (WEL). In: Poschlod P., Borgmann P., Listl D., Reisch C. Zachgo S. (eds.): Handbuch Genbank WEL. Regensburg: HOPPEA Denkschriften der Regensburgischen Botanischen Gesellschaft, 41-69.


 +49 541 969 2840
+49 541 969 2840 szachgo(at)uni-osnabrueck.de
szachgo(at)uni-osnabrueck.de Division of Botany: Barbarastr. 11, D-49076 Osnabrück
Division of Botany: Barbarastr. 11, D-49076 Osnabrück Botanical Garden: Albrechtstr. 11, D-49076 Osnabrück
Botanical Garden: Albrechtstr. 11, D-49076 Osnabrück